
Models of Markov processes are used in a wide variety of applications, from daily stock prices to the positions of genes in a chromosome. Hidden Markov Models (HMM) seek to recover the sequence of states that generated a given set of observed data. This MATLAB function given a sequence, seq, calculates the most likely path through the hidden Markov model specified by transition probability matrix, TRANS,.
Description
[ESTTR,ESTEMIT] = hmmtrain(seq,TRGUESS,EMITGUESS) estimatesthe transition and emission probabilities for a hidden Markov modelusing the Baum-Welch algorithm. seq can be a rowvector containing a single sequence, a matrix with one row per sequence,or a cell array with each cell containing a sequence. TRGUESS and EMITGUESS areinitial estimates of the transition and emission probability matrices. TRGUESS(i,j) isthe estimated probability of transition from state i tostate j. EMITGUESS(i,k) is theestimated probability that symbol k is emittedfrom state i. hmmtrain(..,'Algorithm',algorithm) specifiesthe training algorithm. algorithm can beeither 'BaumWelch' or 'Viterbi'.The default algorithm is 'BaumWelch'.hmmtrain(..,'Symbols',SYMBOLS) specifies the symbols that are emitted. SYMBOLS can be a numeric array, a string array, or a cell array of the names of the symbols. The default symbols are integers 1 through N, where N is the number of possible emissions.Gemini tv chakravakam serial today episode.
hmmtrain(..,'Tolerance',tol) specifiesthe tolerance used for testing convergence of the iterative estimationprocess. The default tolerance is 1e-4.hmmtrain(..,'Maxiterations',maxiter) specifiesthe maximum number of iterations for the estimation process. The defaultmaximum is 100.hmmtrain(..,'Verbose',true) returns thestatus of the algorithm at each iteration.hmmtrain(..,'Pseudoemissions',PSEUDOE) specifiespseudocount emission values for the Viterbi training algorithm. Usethis argument to avoid zero probability estimates for emissions withvery low probability that might not be represented in the sample sequence. PSEUDOE shouldbe a matrix of size m-by-n,where m is the number of states in the hidden Markovmodel and n is the number of possible emissions.If the i→k emission doesnot occur in seq, you can set PSEUDOE(i,k) tobe a positive number representing an estimate of the expected numberof such emissions in the sequence seq.hmmtrain(..,'Pseudotransitions',PSEUDOTR) specifiespseudocount transition values for the Viterbi training algorithm.Use this argument to avoid zero probability estimates for transitionswith very low probability that might not be represented in the samplesequence. PSEUDOTR should be a matrix of size m-by-m,where m is the number of states in the hidden Markovmodel. If the i→j transitiondoes not occur in states, you can set PSEUDOTR(i,j) tobe a positive number representing an estimate of the expected numberof such transitions in the sequence states.If you know the states corresponding to the sequences, use
hmmestimate toestimate the model parameters.
Tolerance
The input argument '
tolerance' controls howmany steps the hmmtrain algorithm executes beforethe function returns an answer. The algorithm terminates when allof the following three quantities are less than the value that youspecify for tolerance:- The log likelihood that the input sequence
seqisgenerated by the currently estimated values of the transition andemission matrices - The change in the norm of the transition matrix, normalizedby the size of the matrix
- The change in the norm of the emission matrix, normalizedby the size of the matrix
The default value of
'tolerance' is 1e-6.Increasing the tolerance decreases the number of steps the hmmtrain algorithmexecutes before it terminates.maxiterations
The maximum number of iterations,
'maxiterations',controls the maximum number of steps the algorithm executes beforeit terminates. If the algorithm executes maxiter iterationsbefore reaching the specified tolerance, the algorithm terminatesand the function returns a warning. If this occurs, you can increasethe value of 'maxiterations' to make the algorithmreach the desired tolerance before terminating.Published online 2011 Dec 21. doi: 10.6026/97320630007418
PMID: 22347785
This article has been cited by other articles in PMC.
Abstract
Since membranous proteins play a key role in drug targeting therefore transmembrane proteins prediction is active andchallenging area of biological sciences. Location based prediction of transmembrane proteins are significant for functionalannotation of protein sequences. Hidden markov model based method was widely applied for transmembrane topologyprediction. Here we have presented a revised and a better understanding model than an existing one for transmembrane proteinprediction. Scripting on MATLAB was built and compiled for parameter estimation of model and applied this model on amino acidsequence to know the transmembrane and its adjacent locations. Estimated model of transmembrane topology was based onTMHMM model architecture. Only 7 super states are defined in the given dataset, which were converted to 96 states on the basis oftheir length in sequence. Accuracy of the prediction of model was observed about 74 %, is a good enough in the area oftransmembrane topology prediction. Therefore we have concluded the hidden markov model plays crucial role in transmembranehelices prediction on MATLAB platform and it could also be useful for drug discovery strategy.
Availability
The database is available for free at [email protected]@bhu.ac.in
Keywords: Hidden Markov Model, Transmembrane Proteins, MATLAB
Background
Accurate predictive success of transmembrane proteins byapplying hidden markov model [HMM] is frequently used inbiological research. This is fully machine learning approach inwhich genome structure and proteins topology prediction arethe fascinating and most demanding subject in bioinformatics.The body of a HMM is closely compatible to the biologicalentities which is being simulated by the model. Looking thefeasibility of HMM by going through the recent researcharticles, is totally statistical approach that compiled by the set ofstates which have potentially able to emit symbols on the basisof probability [1,]. These states are estimated by modelparameters. Model parameters consist of three probabilities i.e.initial probability, transition probability and emissionprobability.
In the context of accurate prediction method of transmembranetopology many workers had obtained results with more thanfive helices were predicted at a significantly lower accuracythan proteins with five or fewer and in addition the estimationof the standard procedure to resolve the prior work andpresented novel trends that may impact the analysis of entireproteomes []. Furthermore, some workers were addressed amethod to reduce the number of false positives, i.e., proteinsfalsely predicted with membrane helices []. Previously theworkers had worked on the effectiveness of modelregularization, dynamic model modification and optimizationstrategies of model and validated through experimentally [].Several workers were developed five different predictionmethod, these are TMMOD [], PHD, HMMTOP, MEMSAT,and TOPPRED for comparatively better result intransmembrane topology prediction [5]. Although theprediction of transmembrane helices can be analyzed with goodenough score plot but this idea was not successful for otherhelices. Recently the hydrophobic property of transmembranehelices was used for the prediction []. But after some time onefeature was taken into account i.e. abundance of positive chargeresidues [,]. Due to better incorporation of helix length,compositional bias and grammar constraints, HMM is suitablefor the prediction of transmembrane helices. Helical membraneproteins are specified a “grammar”, in which cytoplasmic andnon-cytoplasmic loops have occurred alternate fashion.Therefore this feature provides the efficient information, evenperforms better result in prediction [,]. TMHMM approachis applied here because these days mostly membrane protein ispredicted through this method and result is more accurate. Forbetter understanding of model we applied here MATLAB.MATLAB is widely used to visualize and analyze the biologicaldata. MATLAB provides a bioinfo tool box in which variousbioinformatics tools are available for technical computing andscripting. This language is a high-level matrix/array languagewith control flow statements, functions, data structures,input/output, and object-oriented programming features [9]. Itwas experienced that MATLAB shows better feasibility withHMM rather than other bio-statistical packages.
In this work we present our model performance, based onhidden markov model, by taking approach from previousresearch on TMHMM []. We have reviewed the previousTMHMM model architecture which has specialized modeling ofdifferent regions of membrane proteins like inner, middle andouter region []. These regions are collectively divided by 7locations. The states are connected to each other and transitionsmay be possible between adjacent states. Best possibletransition of a state recruits on the basis of high transitionprobability value, known as transition probability and further20 amino acids are emitted by probability distribution of eachstate.
Methodology
Dataset:
We have downloaded two types of dataset from the TMHMMwebsite in which second dataset has 160 transmembranesequences. Most protein pattern of these dataset wasdetermined by experimentally. The dataset is labeled by threemain locations, these are transmembrane helix (m), inside (i)and outside loops (o) on the basis of the existence of deferentamino acids pattern within a transmembrane region. 10-foldcross validation was applied for the validation of model [,].
Method:
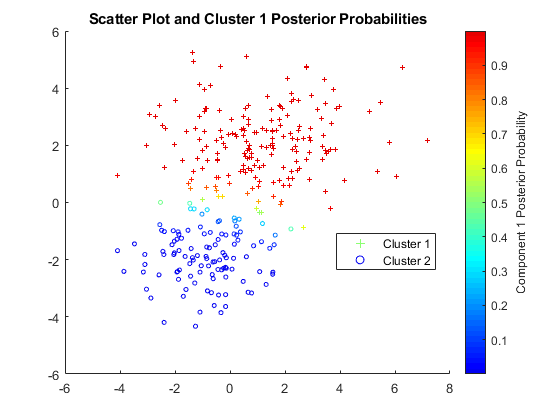
In this work, we have introduced the probabilistic framework ofthe HMM for transmembrane helix prediction. Well knownarchitecture of TMHMM was considered and three mainlocations in dataset were further divided into seven differentsuper states as shown in TMHMM model architecture[12] Figure 1.Conversion from three to seven super states was decided on thebasis of position and length of strings location. In the context ofouter region of transmembrane topology, five residues length ofcap cytoplasmic was used to fix the boundary between helixand loop region. These seven super states were also labeled. Forouter loop, either it was short (S) or long (O) region of noncytoplasmic,was considered the length of 20 residues andsimilarly the loop (L) of cytoplasmic side was counted as 20residues. The cap non-cytoplasmic (N) was considered thelength of 5 residues (Figure 2). The length of helix core (M)region was, determined previously, and considered 25 residues.(Figure 3) The seventh super state, rest part of thetransmembrane architecture was treated as 1 or may be morethan 1. Same text of the state was considered same region. Eachsuper state has an associated probability distribution over the20 amino acids characterizing the variability and pattern ofamino acids in the region. Here the every single position of asuper state was considered as individual state of the model.
Architecture of TMHMM consists of 7 boxes and eachbox indicates the particular region. Same text box treats as sameregion. Here we have denoted each box by symbols to itscorresponding region. Each box may be one or more states inHMM with same parameters.
Expended structure of globular, loop and helix capcytoplasmic region. Possible transitions between the states areshown. Therefore we have considered maximum 10 states forloop on the basis of length of amino acid and same maximum 5states as for helix cap region.
Here representation the detail structure of helix coreand all possible transitions between 25 states.
In this work we have considered the number of states of aregion on the basis of number of amino acid positions []. Forsimilar reasons, a loop's first and last 10 residues are explicitlymodeled, and the each residue corresponds to an individualstate in the model. All other residues in the middle of a loop arecollectively represented by one “globular” state, which has atransition back to itself thus can repeat as many times as theloop length dictates. But in the case of loop apart of other herewe have considered all loop regions for 20 residues includinglong and short loop. Short and long loops were separated hereon the basis of the prediction of adjacent regions by possiblestate transitions. Since long loops outside the membraneappear to have divergent properties than short loops, twoseparate chains of states are introduced, as expressed in figure1.The transmembrane helix model was constructed exactly likeTMHMM [,12], by two cap regions of 5 residues each, including acore region of length 25 residues. Therefore the total length formembrane helices was 35 residues, if we were considered thelength including cap regions, covering the real size rangeobserved for transmembrane domains. The state diagram forthe overall transmembrane topology collectively wasconsidered minimum 96 states by converting these 7 superstates on the basis of length corresponding to its transitions. Theentire states model is thus 20+5+25+5+20+20+1 = 96. Thereforethe IP (Initial probability), STP (State transition probability) andREP (Residue emission probability) matrices were 96×1, 96×96and 96×20 respectively. Each state should be emitted singleamino acids.
HMM Training and Scoring:
In the first stage the aim is to fix the boundaries oftransmembrane helices. Previously many worker weresuggested for boundary correction [] in labeled dataset, like M,I and O. ‘M’ represents membrane region, ‘I’ is for inside region(cytoplasmic) and ‘O’ means outer region (non-cytoplasmic).Therefore we were compiled the MATLAB script for convertingthese 3 labeled dataset sequences to 7 labeled sequences on thebasis of TMHMM model architecture (Figure 1). New modelestimation was done using the relabeled sequences. In Secondstep for estimating the HMM model parameters, posteriorprobability method were applied. Each state of the model hasbeen predicted by forward-backward algorithm, called asposterior probability method. Previously the scoring methoddescribed in the article was implemented using N-bestalgorithm [], was also very effective but here it was foundthat the posterior probability method is more convenient andcompatible for the estimation. This HMM model parameter isestimated in 7 super states and the training was performed byestimating the maximum likelihood []. As usual we have alsoapplied the traditional method (posterior probability) forestimating the HMM and its transmembrane model parameterθ. Here the joint probability of a set of sequence P (x1,…….,xn)being emitted the symbol using a particular state of path [1,]P (x1,x2,……., xn/ θ).
The parameter θ contains all the STP (State TransitionProbability), REP (Residue Emission Probability) and IP (Initialprobability) matrices. Probability of a residue for a particularstate has been evaluated by posterior probability method [,1].Baum-Welch algorithm is standard method for maximumlikelihood estimation of HMMs, in which posterior probabilitieswere performed by utilizing both forward and backwardalgorithms. These algorithms were used to compile the STP andREP matrices. The program was set to 15 iteration time toconverse the values. Initial probability was arbitrary taken forthe initializing the parameter θ calculation. Here we were useddataset sequences for the model estimation and training. Theprocedure used for training a model was the same regardless ofthe scoring method used and in most aspects identical to theprocedure used by Krogh et al. (2001) []. Here, both trainingand scoring was done using the original data set of 160, whichwe have divided it into subset of 16×10.
Validation:
Since 160 dataset have been reported. For ten cross foldvalidation we were divided 160 sequences set into 10 subset.Created 10×16 data and put newly estimated HMM parameteron this 10 subset. In the result, MATLAB calculated the scoresfor each sequences of every subset. We have found the score foreach sequences of all subset. Test set were evaluated by originaltest structure taken from dataset [].
Discussion
Labeled 160 sequences were recruited, measuring the accuracyof model whose topology and locations of transmembranehelices are correctly predicted by TMHMM. The score of eachsequence was calculated by posterior probability method ofHMM for each sequence of 160 dataset. The 16×10 score plot isshowing in Figure 4. Legend of figure shows the 160 stars ofdifferent sequences. Each subset of 16 sets of amino acidsequences possesses 10 sequences. Plot was represented thescores of 16×10 data. Position of a star indicates the score of asequence and the aggregate prediction was calculated byfollowing equation on MATLAB. We were obtainedapproximately 74% accuracy on average. The higher score valueis observed at 98.6. In addition, the 63 times have observed thescore values above 80.
Prediction Score plot for each 10 sequence of 16 set.Dataset sequences were helpful for the validation of model andwe have treated them as a unified set in order to find generalprinciples.
Aggregate prediction = mean (sum (score, 2)/16)
This HMM-based method embodies many conceptual andmethodological aspects of previous methods. The main realitiesare that the model architecture closely to the transmembranepattern and that everything is done in the probabilisticframework of HMMs, so we do not have to develop aspecialised dynamic programming algorithm or posteriorprobability method. Our model prediction is helpful for betterunderstanding of existing model.
Conclusion
All transmembrane sequences for validation of model wereaccessed from TMHMM site indicating that they are allrecruited to the transmembrane helices. Specific transmembranetopology prediction is helpful for protein functional annotation.Along with other documented programming languages,MATLAB was also taken into consideration because theMATLAB performed better compatibility with biologicalentities. Therefore the hidden markov model could performcrucial role in transmembrane helices prediction on MATLABplatform and it could also be useful for drug discovery strategy.Since there are many various statistical approaches are using byseveral workers in biological sciences but HMM is well knownfor higher accuracy result in the area of protein topologyprediction. Meanwhile there should be more work yet to beexplored for higher result accuracy and low level ofredundancy for transmembrane proteins prediction.
Footnotes
Citation:Chaturvedi et al, Bioinformation 7(8): 418- 421 (2011)
References
1. BH Juang, LR Rabiner. Technometri. 1991;33:251.[Google Scholar]
2. PC Chen, et al. Protein Sci. 2002;11:2774.[PMC free article] [PubMed] [Google Scholar]
3. B Rost, et al. Protein Sci. 1996;5:1704.[PMC free article] [PubMed] [Google Scholar]
4. R Hughey, A Krogh. Comput App Biosci. 1996;12:95. [PubMed] [Google Scholar]
5. L Kall, EL Sonnhammer. FEBS Lett. 2002;18:415.[Google Scholar]
6. P Argos, et al. Eur J Biochem. 1982;15:565. [PubMed] [Google Scholar]
7. G Heijne, C Manoil. Protein Eng. 1990;4:109. [PubMed] [Google Scholar]
8. A Krogh, et al. J Mol Biol. 2001;19:567. [PubMed] [Google Scholar]
9. MATLAB technical documentation. Www.mathworks.comRetrieved 2010-06-07. [Google Scholar]
10. EL Sonnhammer, et al. Proc Int Conf Intell Syst Mol Biol. 1998;6:175. [PubMed] [Google Scholar]
11. A Krogh. Proc Int Conf Intell Syst Mol Biol. 1997;5:179. [PubMed] [Google Scholar]
12. R Durbin, et al. Cambridge, UK: Cambridge University Press; 1998. [Google Scholar]
13. G Heijne. EMBO J. 1986;5:3021.[PMC free article] [PubMed] [Google Scholar]
14. RY Robel Kahsay, et al. Bioinformatics. 2005;21:1853. [PubMed] [Google Scholar]
15. G Von Heijne, et al. J Mol Biol. 1992;225:487. [PubMed] [Google Scholar]
16. L Kall, et al. J Mol Biol. 2004;338:1027. [PubMed] [Google Scholar]
Articles from Bioinformation are provided here courtesy of Biomedical Informatics Publishing Group

